

J. Reichert, A, Jabs, P. Slickers, J. Sühnel,
The IMB Jena Image Library of Biological Macromolecules,
Nucleic Acids Research (in press) - 2000 NAR Biological
Database Issue
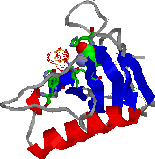
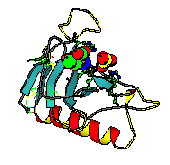
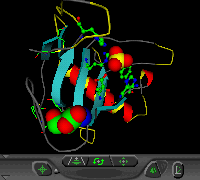
Each automatically generated entry page of the IMB Jena Image Library of Biological Macromolecules contains, at least, one thumbmail image, two vector graphics and one VRML representations, one Rasmol/Chime and one WebMol visualization. In addition, ligands, active sites, and cis-peptide bonds can be visualized separately. Follow this link to see the complete entry page provided by the Image Library for a protein, the Ustilago sphaerogena ribonuclease U2. On this page one can also find links to the websites of molecular graphics programs used. In Figure 1 the various visualization types are displayed.
Figure 1. Visualization types for
automatically generated entry pages (example: Ustilago sphaerogena
ribonuclease U2; PDB code: 1rtu).
[top]
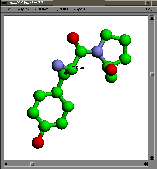
complete structure
(RasMol, GIF and program)complete structure
(Molscript, PDF)complete structure
(WebMol, Java applet)complete structure
(Molscript, VRML)
enlarge image (GIF);
RasMol | Chime linkenlarge image (PDF);
mono | stereoenlarge image (GIF) ;
WebMol linkenlarge image (GIF) ;
VRML
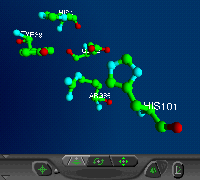
active site:
Tyr39, His41, Glu62, Arg85, His101
(VRML)cis-peptide bond:
Tyr39 , Pro40
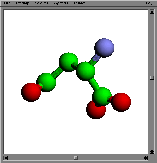
(RasMol)ligand:
aspartyl group
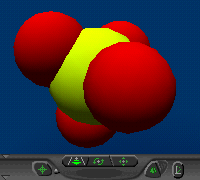
(RasMol)ligand:
sulfate
(VRML)
enlarge image (GIF) ;
VRMLenlarge image (GIF) ;
Rasmol | Chime linkenlarge image (GIF) ;
RasMol | Chime linkenlarge image (GIF) ;
VRML