

J. Reichert, A, Jabs, P. Slickers, J. Sühnel,
The IMB Jena Image Library of Biological Macromolecules,
Nucleic Acids Research (in press) - 2000 NAR Biological
Database Issue
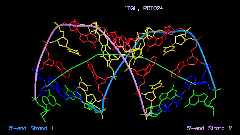
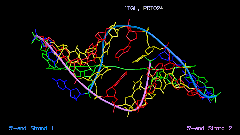
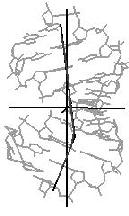
The Image Library includes a tool for the quantitative analysis of helix and bending properties of approximately 750 nucleic acid double helix structures (free or bound to drugs or proteins). It offers both numerical and visual information on the helix geometry. As shown in Figure 3 for a nucleic acid-protein complex the orientation of the nucleic acid part is not just taken from the PDB or NDB file. Rather, the coordinate axes are aligned to the principal axes of inertia of the helix axis. In this manner the bending features of the nucleic acid helix are clearly shown.
Figure 3. Three orthogonal views of
the nucleic acid part of the complex of human TATA-binding protein with
TATA-sequence DNA (PDB code: 1tgh, NDB code: pdt024).
The helical axis is determined with the CURVES algorithm and
subsequently the following geometrical models are fitted to this axis:
a straight line, a circular line (arc), a kinked line, and a double
kinked line. By means of a goodness-of-fit criterion the most
appropriate model is selected. This information and the geometrical
parameters of the corresponding models (radius of curvature, kink and
twist angle) lead to a comprehensive bending classification. In Figure 4 representative examples for the different
bending types are shown.
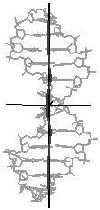
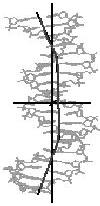
straight line
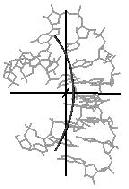
curved line (arc)
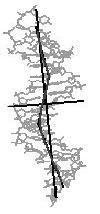
kinked line
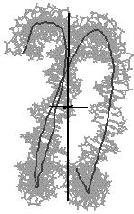
double kinked line
double kinked line
complex line
protein/DNA complex
RNA duplex
DNA/RNA hybrid
protein/DNA complex
DNA duplex
protein/DNA complex
[top]

enlarge image
enlarge image
enlarge image
Figure 4. Helical axis
bending types of nucleic acid duplex structures. The links lead to the
bending analysis pages.
(small twist angle)
(large twist angle)
(Eco RI )
-
-
(Cro)
(HIV-1 kappa B site)
(nucleosome core particle)